snps explained
What is GenoSight? AI Genetic Analysis Explained
GenoSight turns your raw 23andMe, AncestryDNA, or MyHeritage file into an AI-synthesized health report grounded in your full nine-domain health profile.
Sebastian Thorp · May 1, 2026 · 5 min read
In short
GenoSight is an AI-powered genetic analysis platform. You upload the raw genotype file from your 23andMe, AncestryDNA, or MyHeritage test, fill in a nine-domain health profile, and the system returns a personalized health report. Three analysis engines (curated lifestyle SNPs, drug-gene interactions, disease-associated variants) are synthesized by Anthropic Claude into something you can actually act on, plus a chat that lets you ask follow-up questions grounded in your specific findings.
The 30-second version
GenoSight reads your raw DNA file the way a knowledgeable friend with a biology degree might — but at scale, against your specific health context, and with primary-source citations. You bring the raw genotype file (the same one your testing service lets you download). The platform does the rest.
The output isn't a generic "you have variant X" lookup table. It's a synthesized report that asks questions like "given this person's MTHFR genotype, current B-complex stack, and family history of cardiovascular disease, what's actually worth changing?"
The problem GenoSight is built around
Consumer DNA tests give you two things: a polished but shallow report (23andMe's wellness traits, AncestryDNA's genealogy summaries) and a several-hundred-thousand-row raw genotype file you can download. The space between them is where most people get stuck.
Existing options to bridge that gap fall into three buckets:
- Variant lookups (Promethease, SNPedia-driven tools) return a long list of "you have this variant, here's what a study said." Comprehensive but unfiltered — most of it is noise for any given person.
- Editorial libraries explain what each variant means in long-form articles. Excellent depth, but you have to map your own genotype against each article one variant at a time.
- AI-driven platforms are the closest analog to what we built. Functional differences come down to citation depth and how much of your full health context the synthesis actually uses.
The gap GenoSight is built for: take your raw file, take your full health context, and synthesize one cohesive report — with primary-source citations and a chat that lets you ask the follow-up questions every static report leaves unanswered.
How the analysis works
Three analysis engines run against your raw file:
- Curated lifestyle SNPs. A hand-picked set of 79 variants across 16 categories (sleep, caffeine metabolism, sodium sensitivity, methylation, vitamin D conversion, and others), chosen because they have actionable, lifestyle-modifiable downstream effects.
- Pharmacogenomics (PharmGKB Levels 1A–2B). Drug-gene interactions where the evidence is strong enough to inform prescribing decisions — SSRIs and CYP2C19, warfarin and VKORC1/CYP2C9, clopidogrel and CYP2C19, and others.
- Disease-associated variants (ClinVar). 341,375 ClinVar SNPs scanned against your file, filtered to gold-star confidence and true SNPs (indels excluded).
Then Anthropic Claude synthesizes the engine outputs against your health profile. (For the full five-stage pipeline — extraction, engine matching, context fold-in, synthesis, citation — see how AI reads your DNA.) This is the step that turns "you have rs1801133 (CT)" into "given that you're already on B-complex and your homocysteine was 11 µmol/L last test, your current methylated-folate dose is appropriate; recheck homocysteine in 12 weeks before changing anything."
The nine health profile domains
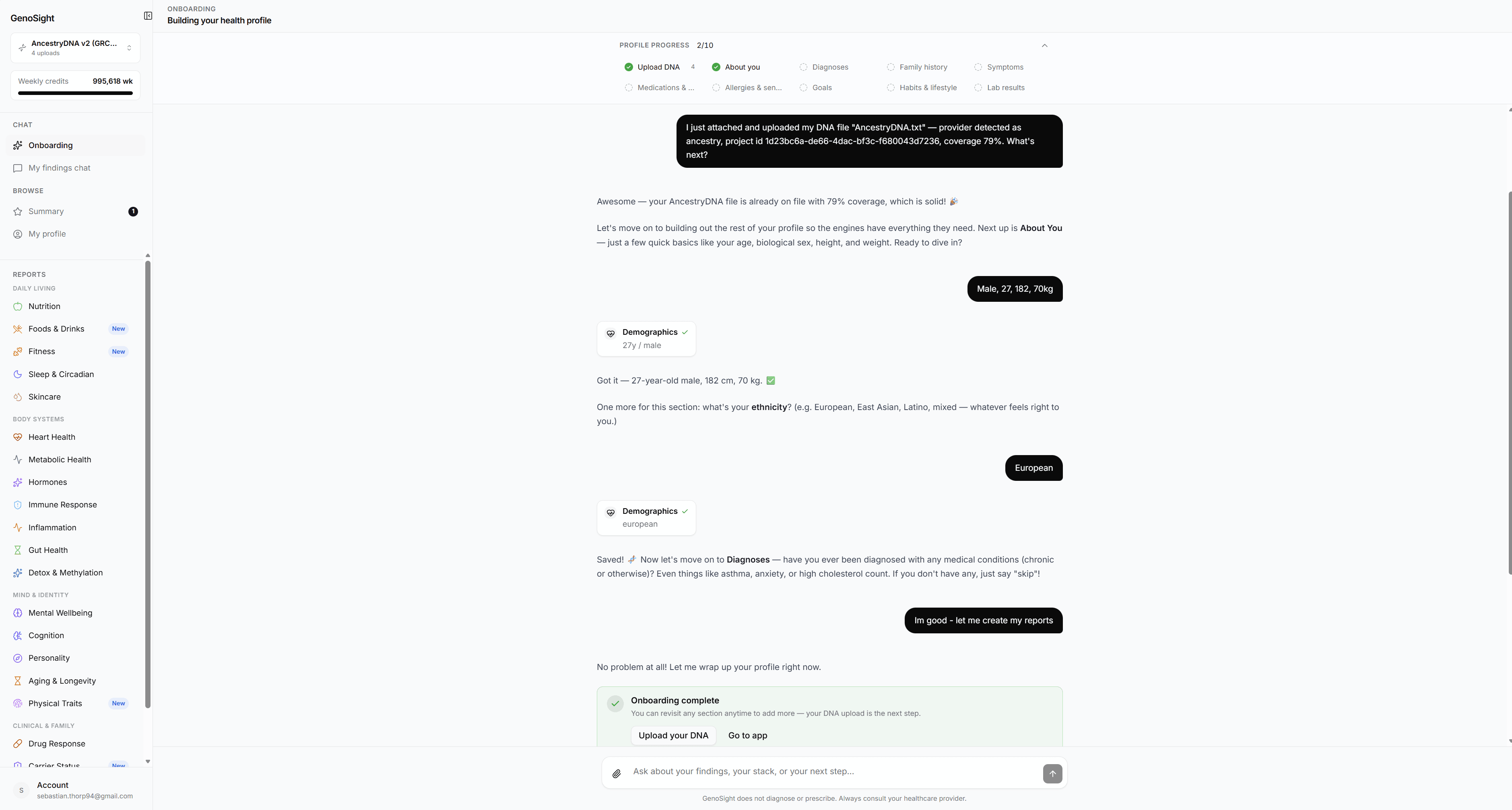
Synthesis quality depends on context. The onboarding chat captures nine domains:
| Domain | What we ask about |
|---|---|
| Demographics | Age, sex, ancestry — affects allele frequency interpretation |
| Diagnoses | Current and past medical conditions |
| Family history | First- and second-degree relatives' diagnoses |
| Symptoms | Current concerns you want the report to address |
| Current stack | Medications and supplements you're taking |
| Allergies | Drug, food, and environmental sensitivities |
| Habits | Diet, sleep, exercise, alcohol, caffeine patterns |
| Goals | What you want to optimize for |
| Recent labs | Numerical values from blood panels, when available |
A C677T homozygous result reads completely differently for someone with no symptoms and a clean lipid panel than for someone with low B12, elevated homocysteine, and treatment-resistant depression. Without that context, every report would sound the same. (Walk through each domain in detail.)
Findings-grounded chat
After the report renders, you can ask follow-ups:
- "Why methylcobalamin and not cyanocobalamin in my case?"
- "What dose makes sense given that I'm already on SAM-e?"
- "Should I retest homocysteine before or after starting?"
- "What does ClinVar's two-star confidence actually mean for this variant?"
Each answer is grounded in your specific findings, not a generic FAQ. The chat is read-only by default; the one tool that can change anything (propose_stack_change) requires explicit confirmation before it touches your supplement or medication list.
What you get back
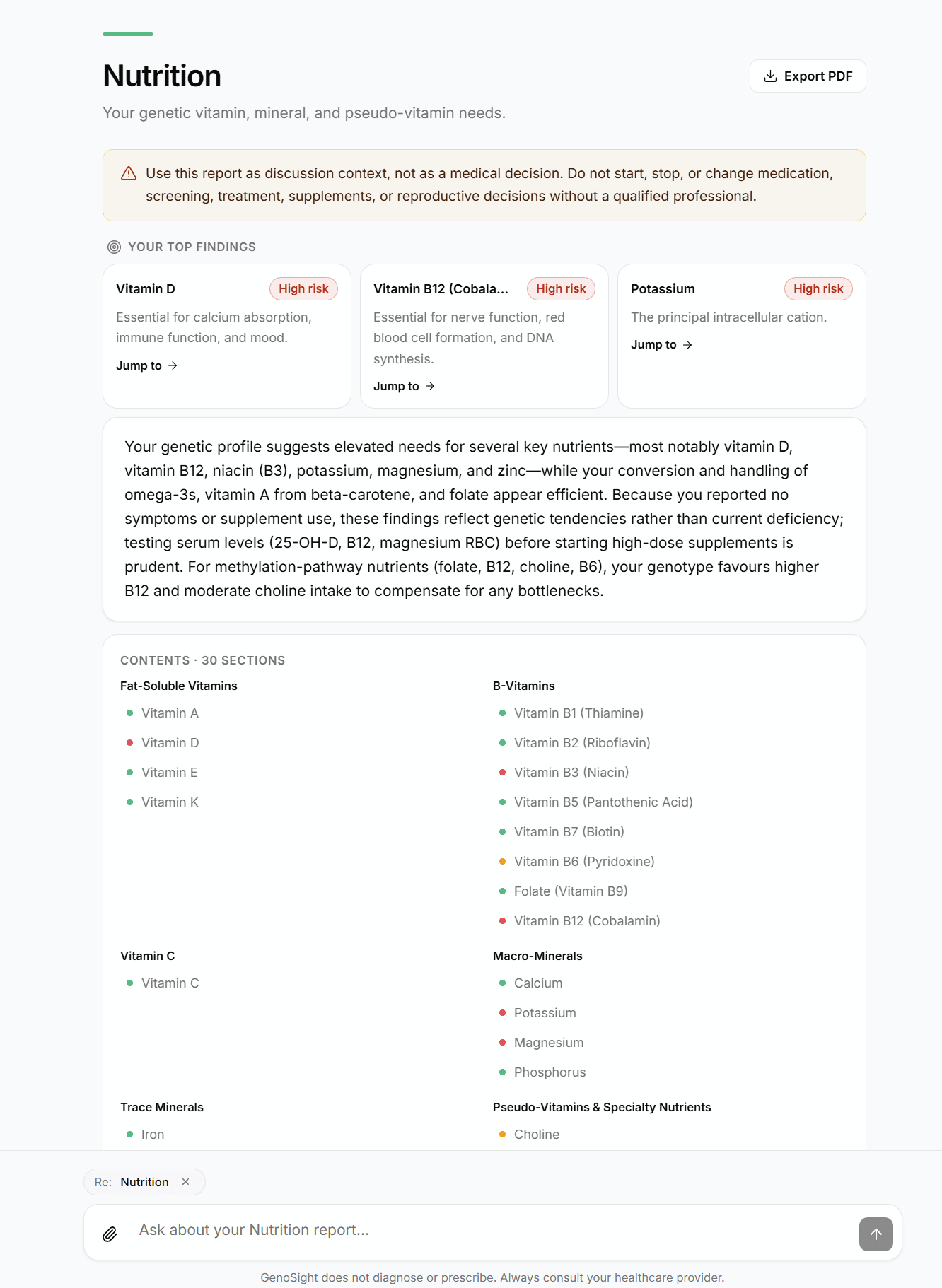
- An on-screen report you can re-read and search
- A PDF version emailed to you (clean enough to hand to a clinician)
- The chat, available indefinitely against your finalized findings
Average report generation runs under 60 seconds.
What GenoSight is not
This part matters as much as the rest. GenoSight is not a medical device, not a diagnostic test, not a substitute for clinical-grade genetic testing or a genetic counselor. Consumer-genomics services use genotyping arrays that cover several hundred thousand positions, not the roughly three billion in your full genome. That's enough for the variants in our analysis engines, but it isn't whole-genome sequencing.
We also won't claim that any variant will cause any outcome. The vocabulary throughout the platform is "may influence" and "is associated with" — never "causes." Variant interpretation evolves as new studies publish; reports are point-in-time and meant to be re-run as the evidence base changes.
Privacy posture
Your raw genotype file is encrypted at rest. Database access is restricted by row-level security so each user can only read their own data. Critically: the raw file itself is never sent to the LLM. The analysis engines extract a small set of relevant variant calls; only those calls and your health profile go to Claude for synthesis. (Full privacy posture here.)
Try GenoSight free
New accounts get 250 signup credits — enough for one deeper report or two lighter ones. No card required.
Medical disclaimer
GenoSight provides educational information about your genetic data. It is not a medical diagnosis, treatment, or cure. Always consult your healthcare provider before making decisions based on this information. Variant interpretation evolves; recheck periodically.
Key takeaways
- GenoSight reads the raw genotype file you download from 23andMe, AncestryDNA, or MyHeritage and returns an AI-synthesized health report — not a generic variant lookup.
- Three analysis engines run against your file: 79 curated lifestyle SNPs, PharmGKB Level 1A–2B drug-gene interactions, and 341,375 ClinVar disease variants.
- Synthesis is contextual: the report reasons about YOUR genotype against your nine-domain health profile (diagnoses, family history, current stack, labs, and others), not a generic genome.
- A findings-grounded chat lets you ask follow-up questions; answers cite the specific variants and evidence behind each recommendation.
- This is educational, not diagnostic. Consumer genotyping arrays cover several hundred thousand variants — useful, but not a substitute for clinical-grade testing or a genetic counselor.
Sources
- ClinVar — NCBI archive of variants of clinical significance — https://www.ncbi.nlm.nih.gov/clinvar/
- PharmGKB — Pharmacogenomics Knowledgebase, Stanford University — https://www.pharmgkb.org/
- GWAS Catalog — EBI catalog of published genome-wide association studies — https://www.ebi.ac.uk/gwas/
- Anthropic Claude — model documentation and safety posture — https://docs.anthropic.com/