snps explained
How AI Reads Your DNA: Inside the Synthesis Pipeline
AI doesn't replace genetic science — it removes the bottleneck between raw variant calls and personal context. Here's exactly how an AI genetic report is built.
Sebastian Thorp · May 1, 2026 · 6 min read

In short
AI doesn't read your DNA the way a human geneticist does — it doesn't need to. The genetic-science work (which variants matter, what they do, how confident the evidence is) is already encoded in curated databases like ClinVar, PharmGKB, and the GWAS Catalog. What AI adds is the synthesis step: matching your specific variant calls against your specific health context and producing one coherent report instead of a thousand isolated lookups. This walkthrough explains the five-stage pipeline GenoSight uses, what the AI is and isn't doing, and where the guardrails sit.
Why "AI genetic analysis" sounds vague — and what it actually means
The phrase gets used loosely. Some products use AI for one narrow step (a chatbot wrapping a static report). Others use it end-to-end (synthesis, prioritization, follow-up Q&A). Most don't tell you which.
The honest framing: AI is most useful for the part of genetic interpretation that isn't a science problem — it's a context problem. The science (this variant is associated with this trait at this evidence level) is settled in curated databases. The bottleneck is mapping that evidence against your situation: your medications, your symptoms, your family history, your goals.
That mapping used to require either a geneticist or a lot of patient self-research. AI removes the bottleneck.
The five-stage pipeline
Here's exactly how a GenoSight report is built from your raw genotype file. Each stage is auditable and has a different failure mode.
Stage 1 — Extraction
The raw file from 23andMe, AncestryDNA, or MyHeritage is a tab-separated list of variant calls — typically several hundred thousand rows of rsID, chromosome, position, genotype. Stage 1 parses the file, normalizes formats across the three providers, and handles strand-orientation differences (a variant reported as AG in one format may be CT in another at the same position).
No AI in this stage. It's pure parsing — deterministic, testable, and the place where most "DNA analysis" tools either succeed or silently produce garbage downstream.
Stage 2 — Engine matching
Three analysis engines run in parallel against the parsed file:
- Lifestyle SNP engine. Looks up your calls against a curated catalog of 79 variants across 16 categories (sleep, caffeine, sodium, methylation, vitamin D conversion, and more). For each match, it returns the genotype and the cited effect.
- Pharmacogenomics engine (PharmGKB Levels 1A–2B). Matches your calls against drug-gene interaction tables — SSRIs and CYP2C19, warfarin and VKORC1, and others. Only Levels 1A through 2B are surfaced (the highest evidence tiers).
- Disease-variant engine (ClinVar). Scans 341,375 ClinVar SNPs, filtered to gold-star confidence. Indels are excluded (consumer arrays don't reliably call them).
Engine outputs are structured findings: variant + genotype + effect + evidence level + citation. Still no AI — these are deterministic database lookups. (Why these four sources.)
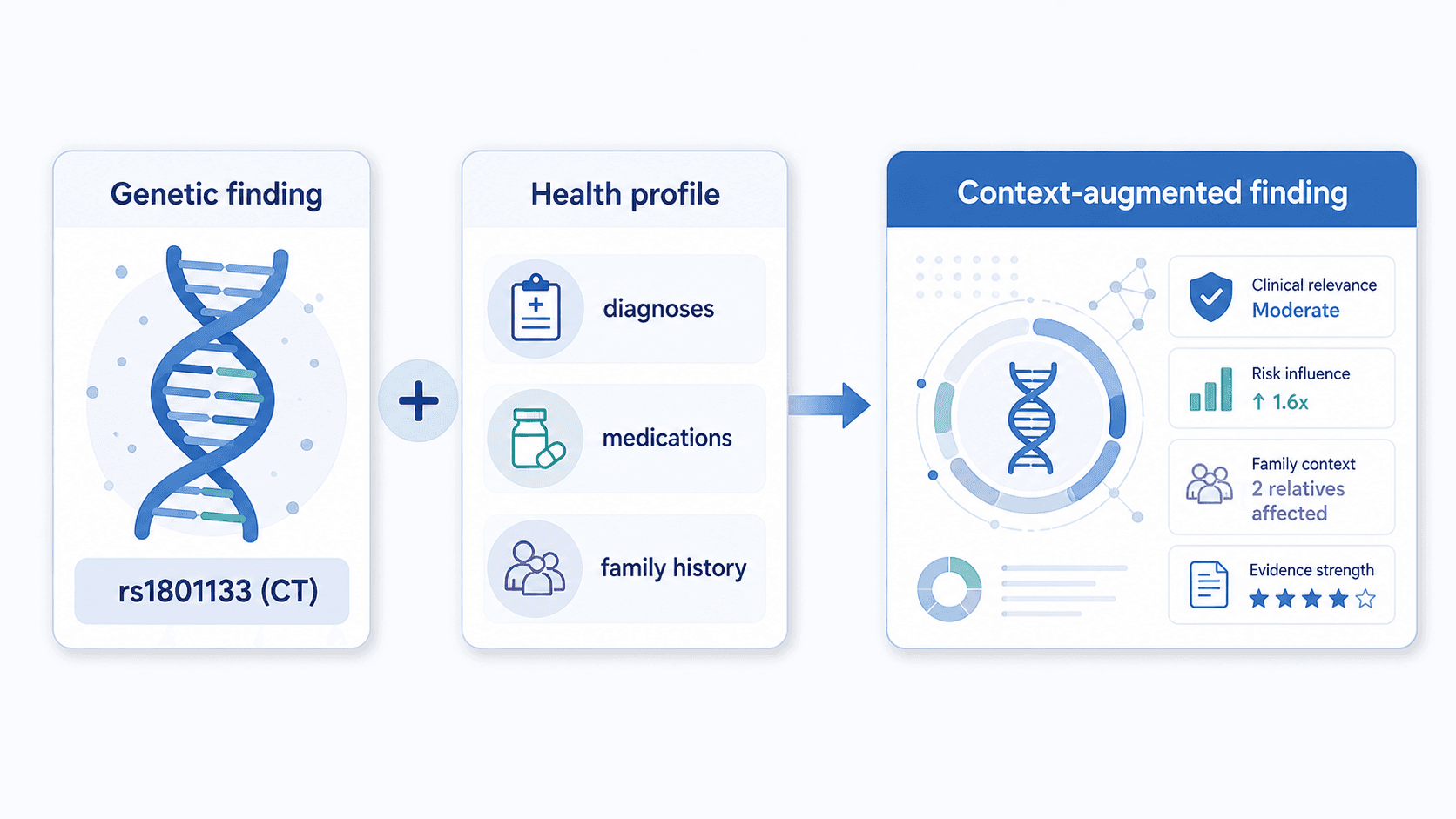
Stage 3 — Context fold-in
This is where your nine-domain health profile enters the pipeline. The engine findings get joined against your demographics, diagnoses, family history, symptoms, current medications and supplements, allergies, habits, goals, and recent lab values.
The output is a structured "context-augmented findings" object — each genetic finding now carries the relevant context that should inform how it's interpreted.
A C677T heterozygous result with a clean homocysteine lab and no symptoms is a different finding than the same variant in someone with elevated homocysteine and treatment-resistant depression. Stage 3 is where that distinction gets made explicit, before AI sees anything.
Stage 4 — Synthesis
This is the AI step. The context-augmented findings get sent to Anthropic Claude, with a prompt that asks: given these findings and this user context, produce a structured report covering an executive summary, prioritized findings, supplement and lifestyle recommendations, drug-gene interactions, carrier status notes, and a monitoring schedule.
A few specific design choices matter here:
- The raw genotype file is never sent to the LLM. Only the structured findings (variant + genotype + effect) and the health profile go to Claude. The raw file stays in encrypted storage. (Full privacy detail.)
- The model is instructed to use only the supplied findings. It cannot invent variants, evidence levels, or citations. Hallucinations get caught by post-synthesis validators that re-check every cited variant against the engine outputs.
- The vocabulary is constrained. "May influence," "is associated with," "evidence suggests" — never "causes," "guarantees," or "treats." This is enforced both at the prompt level and by a validator that flags forbidden phrases before the report is delivered.
Stage 5 — Citation and validation
Every claim in the synthesized report is mapped back to its underlying engine finding. Any sentence that makes a factual claim without a corresponding citation is flagged. Reports that fail this check go to an internal review queue rather than being delivered.
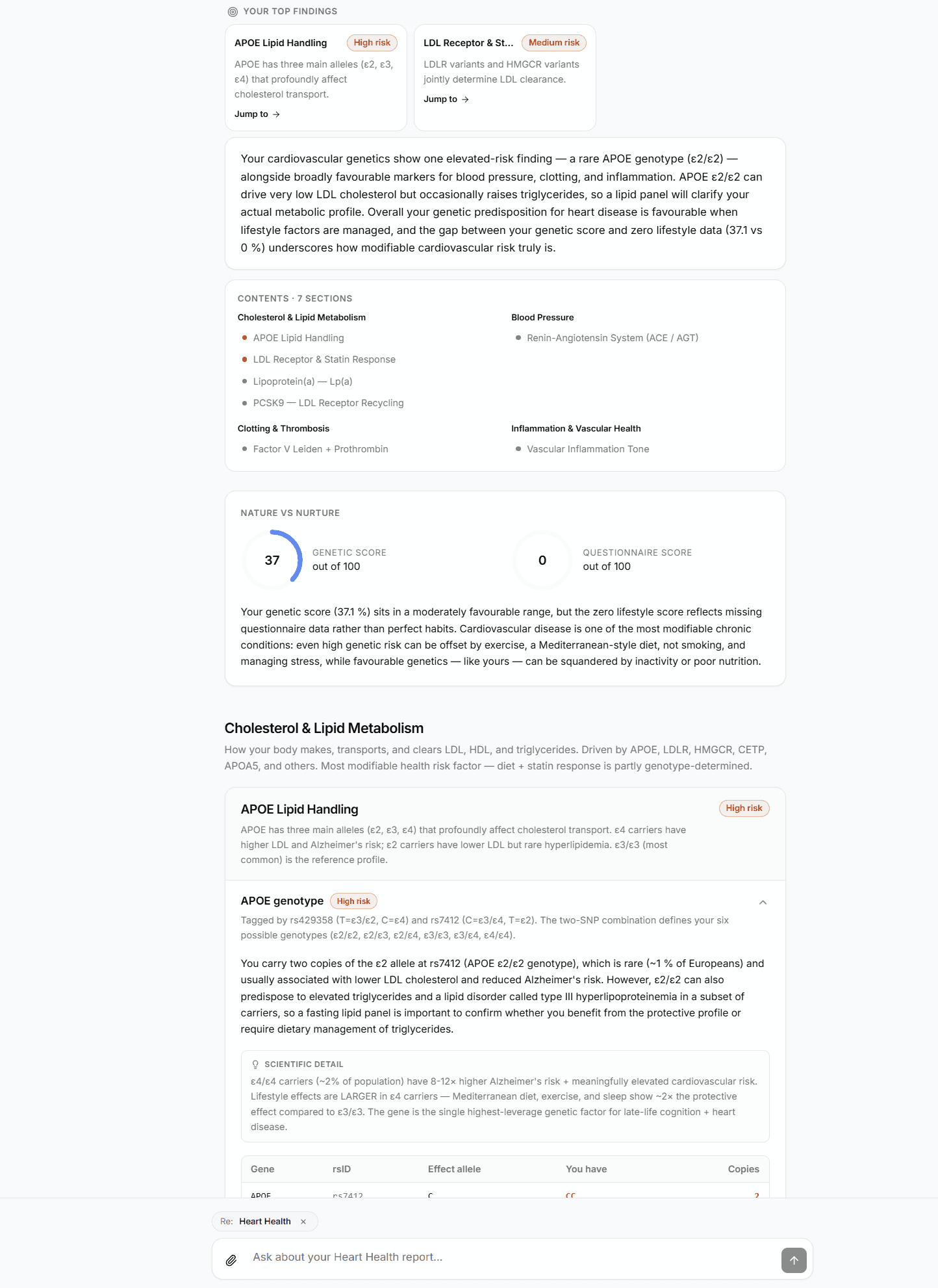
The output is the report you read on screen and the PDF that gets emailed to you.
What the AI does — and doesn't — do
| The AI does | The AI doesn't |
|---|---|
| Synthesize structured findings into a coherent report | Decide which variants matter (that's the engine catalog) |
| Prioritize findings against your specific health context | Invent variants, genotypes, or evidence |
| Write follow-up answers grounded in your specific findings | Diagnose conditions or prescribe treatments |
| Use plain language while preserving precision | Replace your clinician |
| Flag interactions between findings (e.g., MTHFR + COMT) | Modify your stack without explicit confirmation |
The chat interface is read-only by default. The single tool that can propose a change to your supplement or medication stack (propose_stack_change) requires explicit user confirmation in-flow before anything is recorded.
The follow-up question matters more than the report
A static report — even a good one — can't anticipate every question a reader will have. The chat is where AI earns its keep:
- "Why methylcobalamin specifically? My doctor asked."
- "How does my CYP2C19 status interact with the SSRI my psychiatrist suggested?"
- "What does ClinVar's two-star confidence actually mean for this variant?"
- "If I retest homocysteine in 8 weeks, what value would suggest the protocol is working?"
Each answer is grounded in your specific findings, not a generic FAQ. Each chat message costs 1 credit — built into the credit packs across all paid plans.
Why Anthropic Claude, specifically
Three reasons. First, Claude has strong scientific reasoning and a comparatively low hallucination rate on structured-input tasks like this one. Second, Claude's prompt-caching and citation tooling make it efficient at large structured-context synthesis. Third, the safety posture and refusal patterns align with the constraints YMYL content requires — Claude won't be cajoled into making diagnostic claims it shouldn't.
That said: the model is one component. The engines, the context fold-in, the validators, and the citation re-check are equally load-bearing. Swapping the model wouldn't change the architecture.
Try GenoSight free
Upload your raw DNA file and see the synthesis output for yourself. 250 signup credits, no card.
Medical disclaimer
GenoSight provides educational information about your genetic data. It is not a medical diagnosis, treatment, or cure. Always consult your healthcare provider before making decisions based on this information. Variant interpretation evolves; recheck periodically.
Key takeaways
- AI doesn't replace genetic science — it removes the bottleneck between raw variant calls and personal context. The science lives in curated databases (ClinVar, PharmGKB, GWAS Catalog); the AI does synthesis.
- GenoSight's pipeline runs in five stages: extraction, engine matching, context fold-in, AI synthesis, and citation validation. Only stage four uses an LLM.
- The raw genotype file is never sent to the LLM — only structured engine findings and your health profile.
- The model is instructed to use only the supplied findings; post-synthesis validators check every claim against the underlying citations and reject hallucinations.
- The follow-up chat is where AI earns its keep — generic reports can't anticipate every question, but findings-grounded answers can.