snps explained
DNA Tests with Raw Data Export: Which Allow Re-analysis
Not every DNA test lets you download and re-analyze your raw file. Here is which services allow it, what file formats they produce, and how to check yours for compatibility.
Sebastian Thorp · May 1, 2026 · 5 min read
In short
Not every consumer DNA test lets you download and re-analyze your raw data. The big three — 23andMe, AncestryDNA, MyHeritage — do. Several other services either don't offer raw export, or produce a file format that's not compatible with most third-party analysis tools. This guide covers which tests support raw data export, what each format looks like, how to download yours, and what to do if your test doesn't support it.
Which tests support raw data export
The short version, then the detail:
| Test | Raw download | File format | Tool-compatible |
|---|---|---|---|
| 23andMe | ✅ | Tab-separated .txt (or zipped) | ✅ |
| AncestryDNA | ✅ | Tab-separated .txt (zipped) | ✅ |
| MyHeritage | ✅ | Tab-separated .csv (zipped) | ✅ |
| Family Tree DNA | ✅ | Tab-separated .csv | ⚠️ Varies by tool |
| LivingDNA | ✅ | Tab-separated | ⚠️ Often yes |
| Nebula Genomics | ✅ | VCF / FASTQ / BAM (whole genome) | ❌ Different format — WGS not accepted by most consumer tools |
| Color Genomics | ⚠️ Limited | Clinical-grade report; no consumer raw export | ❌ |
| Invitae | ⚠️ Limited | Clinical-grade VCF on request | ❌ Clinical tools only |
| AncestryHealth (legacy) | ❌ Discontinued | Same as AncestryDNA when it existed | ✅ If you saved your file pre-shutdown |
✅ Yes · ⚠️ Partial / conditional · ❌ No
The big three (23andMe, AncestryDNA, MyHeritage) all produce similar file structures: a list of variant calls in a tab-separated text format, with each row containing rsID, chromosome, position, and genotype. Most third-party analysis tools accept all three.
How to check whether your raw file is compatible
A typical compatible file looks like this when you open it in a text editor (the column headers vary slightly between providers):
# rsid chromosome position genotype
rs548049170 1 69869 TT
rs9283150 1 565508 AA
rs116587930 1 727841 GG
If you open your download and see roughly that structure (one row per variant, four columns), it's a standard genotyping-array export and most analysis tools will accept it.
If you see something else — a long XML document, a binary file, or a single PDF report — it's not the kind of raw export that gets re-analyzed by most consumer tools.
How to download your raw file from each service
Each service buries the export option slightly differently, but the pattern is broadly:
- Account settings → privacy or data section → download raw data
- Email confirmation, sometimes with 2FA, before the download is generated
- The file arrives as a download link or attachment
Specific service-by-service walkthroughs change as UIs are updated. For the most current step-by-step instructions, the testing service's own help center is the most reliable source.
A practical tip regardless of service: download the file to a known location and keep a backup. Some services revoke download links after a few days; others let you re-download anytime. The file itself is small (a few megabytes compressed) and reading it costs you nothing later.
Why Nebula's format is different
Nebula sells whole-genome sequencing, not genotyping. Their raw output is a VCF (or sometimes FASTQ/BAM) file that covers far more positions and is structured differently from a 23andMe-style genotyping export.
This is a difference in data depth, not just format. A Nebula 30x WGS file contains data on positions a consumer array doesn't read — including many of the rare variants and structural changes consumer arrays miss. The trade-off: most consumer-grade analysis tools (including GenoSight) don't accept WGS files. Tools that do exist tend to be more research- or clinical-focused.
Important update: Nebula Genomics shut down its consumer service in February 2025. If you previously sequenced with them, your file is still yours — but new sequencing is no longer available through them. Sequencing.com is the closest active replacement for whole-genome work. (How GenoSight compares to Nebula and others.)
What if your test doesn't allow raw export
Three options:
- Re-test with a service that does allow export. 23andMe, AncestryDNA, and MyHeritage all run sales periodically. The cost of one of these is usually $50–100 with a few weeks of turnaround.
- Check whether your service offers an export workaround. Some clinical-grade tests will release a structured report (or even a VCF) on request — though the format may not be consumer-tool compatible without conversion.
- For PGx specifically, ask your clinician. Clinical-grade pharmacogenomic panels exist (Genomind, GeneSight, others) and produce reports designed for clinical workflow. They cost more than re-testing with a consumer service, but they cover variants consumer arrays miss.
Other compatibility notes
A few situations that come up:
- 23andMe Health + Ancestry vs Ancestry-only. Both produce the same raw genotype file. The difference is which reports 23andMe gives you on top of the file — the underlying data is the same.
- AncestryDNA vs AncestryHealth. AncestryHealth was discontinued; if you have a download from when it was available, it's the same underlying genotype file as AncestryDNA. New tests with Ancestry only produce the AncestryDNA export.
- Older test versions. All three major services have changed their array chip versions over the years. Older files have slightly different SNP coverage than newer ones. Most analysis tools accommodate this; a small number of variants may not be present in older exports.
- Different ancestry tests over time. If you've taken multiple tests (a 23andMe and an AncestryDNA, for example), each produces its own raw file. You can analyze each independently — the overlap is significant but not 100%.
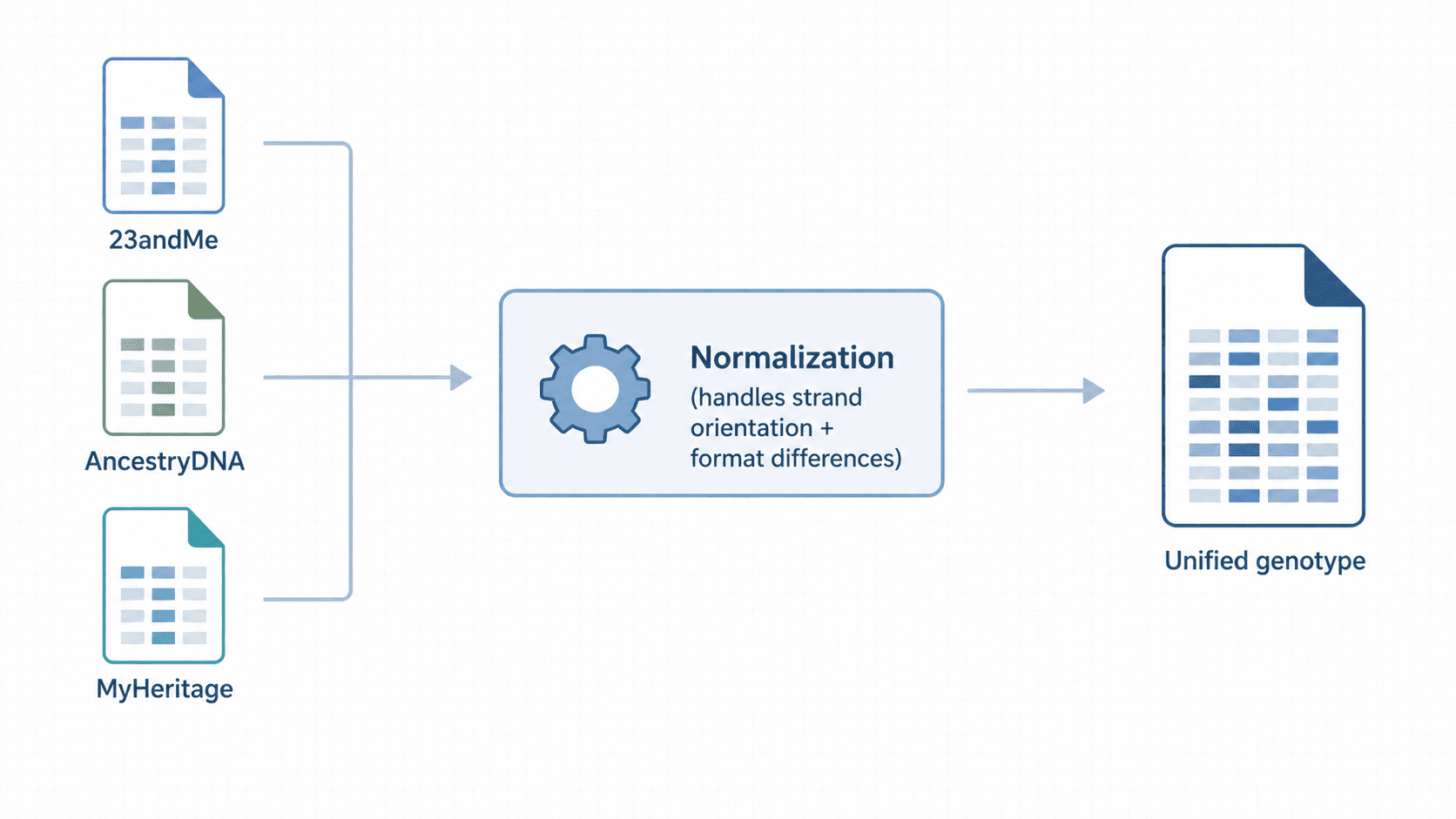
How GenoSight handles file formats
GenoSight accepts raw files from the three main consumer providers — 23andMe, AncestryDNA, and MyHeritage — and handles the format differences automatically. This includes:
- The minor column-naming and ordering differences between providers
- Strand-orientation differences (a variant reported as
AGin one provider's format may beCTin another at the same position; the parser handles bidirectional matching) - Older-version array files that include or exclude specific SNPs
You can upload directly during onboarding without converting the file. If your file format isn't recognized, the upload step explains what went wrong and what to do.
Try GenoSight free
Upload your existing 23andMe, AncestryDNA, or MyHeritage file. 250 signup credits with no card.
Key takeaways
- 23andMe, AncestryDNA, and MyHeritage all let you download your raw data in compatible tab-separated formats — usable by most consumer analysis tools including GenoSight.
- Family Tree DNA and LivingDNA also offer raw exports; compatibility varies by analysis tool.
- Nebula Genomics produces whole-genome sequencing files (VCF/FASTQ/BAM), which are more comprehensive but not accepted by most consumer analysis tools.
- Color Genomics and Invitae are clinical-grade services without consumer-style raw exports — their workflows aren't designed for re-analysis.
- If your test doesn't allow raw export, re-testing with a consumer service is usually $50–100 and produces a file that works across most analysis tools.
Sources
- 23andMe — Genotyping Platform specifications and chip generations — https://customercare.23andme.com/hc/en-us/articles/202904600
- AncestryDNA — Technical overview and FAQ — https://support.ancestry.com/s/article/AncestryDNA-Test
- Nebula Genomics — 2026 status review (consumer service discontinued February 2025) — https://knowyourdna.com/nebula-genomics/
- Sequencing.com — Active whole-genome sequencing platform — https://sequencing.com/