clinical genetics
Limits of Consumer DNA Testing: What It Can't Tell You
Consumer DNA tests look at a few hundred thousand positions, not your full genome. Here is what they reliably catch, what they miss, and when clinical-grade testing matters.
Sebastian Thorp · May 1, 2026 · 6 min read

In short
Consumer genetic tests (23andMe, AncestryDNA, MyHeritage) read your DNA at a few hundred thousand specific positions out of three billion. That's enough for the variants in well-validated lifestyle, pharmacogenomic, and disease catalogs — but it isn't whole-genome sequencing. Some clinically important variants are rare or are insertions/deletions that genotyping arrays can't reliably detect. This guide is honest about what consumer DNA tests can and can't tell you, and where clinical-grade testing remains the right next step.
How consumer DNA tests actually work
23andMe, AncestryDNA, and MyHeritage all use genotyping arrays — small chips with probes for specific positions in the genome. When your sample is processed, the chip checks each probe position and reports the genotype at that position.
A typical consumer array covers between 600,000 and 700,000 positions out of the roughly 3 billion bases in the human genome. That's a tiny fraction — about 0.02% — but it's strategically chosen. The positions are picked to:
- Capture common variants associated with traits and disease (the GWAS-significant SNPs)
- Catch ancestry-informative markers
- Include FDA-authorized clinical reporting variants
- Tag major haplotype blocks (so nearby variants can sometimes be inferred)
That last point matters for imputation — statistical methods that infer untested positions based on the linked variants the array does measure. Imputation is useful for ancestry and for some research applications, but it doesn't reliably rescue rare disease variants.
What consumer arrays reliably catch
For variants the array directly tests, consumer-grade results are good. The accuracy at any given probe position is typically 99%+ when the genotype is unambiguous, comparable to lab-grade testing for those specific positions.
This means consumer arrays handle these well:
- Common variants with established associations — most of the lifestyle SNPs in well-curated catalogs
- Most well-studied pharmacogenomic variants — the major CYP2C19, CYP2D6, VKORC1, CYP2C9 alleles directly tested on the array
- Common carrier variants for the most prevalent recessive conditions — the specific CFTR ΔF508 mutation, the major HBB variants for sickle cell, common HEXA variants
- Common ancestry-informative markers
For the variants the array tests, the limitation isn't accuracy — it's interpretation.
What consumer arrays miss
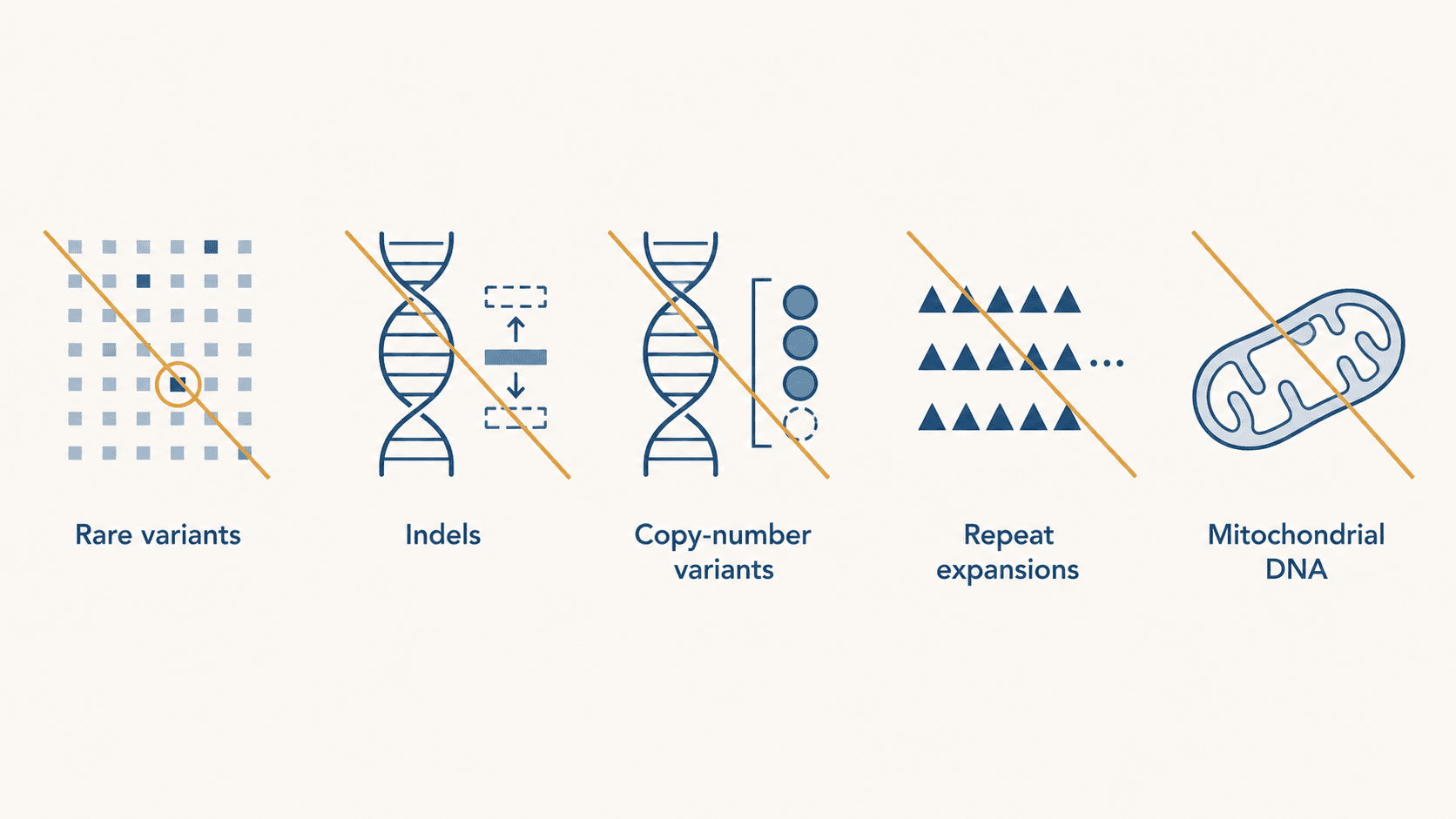
Five categories don't show up reliably in consumer-array data:
Rare variants
If a variant is rare enough that no probe is dedicated to it, the array can't see it. Many disease-causing mutations are rare. Most of the ~2,000 known CFTR variants aren't on consumer arrays. Most rare-cancer-predisposition variants aren't either.
This is the most common misunderstanding about consumer testing. "No variants detected" doesn't mean "no relevant variants exist" — it means "no relevant variants are at the positions we tested."
Insertions and deletions
Most variants the engines reason about are single-nucleotide variants (SNVs) — one base substituted for another. But many disease-causing mutations are insertions (extra bases added) or deletions (bases removed). These are harder for genotyping arrays to call reliably.
GenoSight's ClinVar engine explicitly filters to SNVs for this reason — surfacing indel results from a consumer array would be misleading because the false-call rate is too high.
Copy-number variants and structural changes
CYP2D6 has gene duplications and deletions that meaningfully affect drug metabolism. BRCA1/BRCA2 large deletions exist. These are structural changes, not single-base differences. Consumer arrays don't reliably detect them.
This is one reason clinical-grade PGx panels remain the gold standard for high-stakes prescribing decisions involving CYP2D6: they explicitly call copy-number changes that consumer arrays miss.
Repeat expansions
Conditions like Huntington's disease and fragile X syndrome involve trinucleotide repeat expansions — the same short sequence repeated many times. The number of repeats matters clinically. Consumer arrays measure presence/absence of variants, not repeat counts.
Mitochondrial DNA
Some inherited conditions involve mitochondrial DNA, which is inherited maternally and present in many copies per cell. Most consumer arrays don't comprehensively cover mitochondrial variants.
What this means for your interpretation
A few practical conclusions follow from the above.
A "clean" consumer-array report doesn't rule out genetic risk. It rules out the variants the array directly tested. For most people, that's still informative. For people with strong family history of conditions caused by rare variants, it's not enough.
Negative carrier results need careful framing. "23andMe screened me for cystic fibrosis and I'm not a carrier" really means "23andMe tested for a specific subset of CFTR variants and you don't carry those." For pre-pregnancy decisions, expanded carrier screening from a clinical lab is more comprehensive.
Some PGx interpretation requires clinical-grade panels. Pharmacogenomic interpretation belongs with a prescribing clinician working from a clinical-grade panel — consumer arrays are not a substitute. CYP2D6 in particular often needs clinical-grade panels because of copy-number complexity.
For specific conditions, go directly to clinical-grade testing. Cancer-predisposition panels, cardiomyopathy panels, comprehensive carrier screening — all are clinical-grade workflows for good reasons. Consumer arrays are useful adjuncts; they're not a substitute.
When clinical-grade testing is the right move
Five situations where you should skip the consumer-array workflow and go directly to clinical-grade:
- Strong family history of a specific condition. Hereditary breast/ovarian cancer, Lynch syndrome, hereditary cardiomyopathies, cystic fibrosis with affected family members — these warrant targeted clinical panels.
- Pre-conception expanded carrier screening. Particularly for couples where ancestry, family history, or prior pregnancies suggest higher carrier risk — this is a clinical-grade test ordered through a healthcare provider or genetic counsellor, not something to derive from a consumer array.
- Diagnostic workup for an unexplained condition. When a clinician is investigating a possible genetic cause of symptoms, they'll order targeted clinical-grade testing — not consumer-array re-analysis.
- High-stakes PGx for CYP2D6-metabolized drugs. Clinical-grade panels handle CYP2D6 copy-number complexity that consumer arrays don't.
- Pre-implantation or prenatal testing. These are clinical workflows by definition.
In each of these cases, the consumer-array file (and any analysis built on it, including GenoSight's) is informational input to the conversation, not the answer.
Why we still think consumer-array analysis is worth doing
Despite the limits, three reasons consumer-array analysis remains valuable:
- For most people, the well-validated catalog of lifestyle SNPs the array does cover is genuinely useful — and far more accessible than waiting for a clinical genetic visit. (What GenoSight builds on top of that catalog.)
- The data you already paid for is otherwise idle. A 23andMe or AncestryDNA test you took years ago is sitting in a download folder. Running it through additional analysis costs nothing more.
- It produces structured questions for your clinician — useful for bringing genetic context into a 15-minute appointment — that direct clinical attention to specific decisions.
The honest framing is that consumer-array analysis is a wider lens on a limited dataset. Useful for many things, insufficient for some, never a substitute for clinical-grade testing when the situation warrants it.
Try GenoSight free
GenoSight surfaces what consumer arrays do reliably catch — with explicit caveats about what they don't.
Medical disclaimer
GenoSight provides educational information about your genetic data. It is not a medical diagnosis, treatment, or cure. Always consult your healthcare provider before making decisions based on this information. Variant interpretation evolves; recheck periodically.
Key takeaways
- Consumer DNA tests use genotyping arrays — they read your DNA at 600,000–700,000 specific positions out of about 3 billion in the genome.
- Reliably caught: well-validated common variants, most major PGx alleles, common carrier variants, ancestry markers.
- Missed or unreliable: rare variants, insertions/deletions, copy-number variants (notably CYP2D6 duplications/deletions), repeat expansions, mitochondrial DNA.
- A "clean" consumer-array report doesn't rule out genetic risk — it rules out the variants the array directly tested.
- Clinical-grade testing remains the right move for strong family history, expanded carrier screening, diagnostic workups, high-stakes CYP2D6 PGx, and pre-implantation or prenatal contexts.
Sources
- 23andMe — Genotyping Platform technical specifications — https://customercare.23andme.com/hc/en-us/articles/202904600
- NCBI GeneReviews — Huntington Disease (HTT trinucleotide repeat) — https://www.ncbi.nlm.nih.gov/books/NBK1305/
- PharmGKB — CYP2D6 gene overview and copy-number complexity — https://www.pharmgkb.org/page/cyp2d6
- ACOG Committee Opinion 690 — Carrier Screening for Genetic Conditions — https://www.acog.org/clinical/clinical-guidance/committee-opinion/articles/2017/03/carrier-screening-for-genetic-conditions