clinical genetics
GenoSight Data Sources: ClinVar, PharmGKB, GWAS, PubMed
Every GenoSight finding traces back to a primary source. Here is what ClinVar, GWAS Catalog, PharmGKB, and PubMed are — and why the source choice matters for trust.
Sebastian Thorp · May 1, 2026 · 6 min read
In short
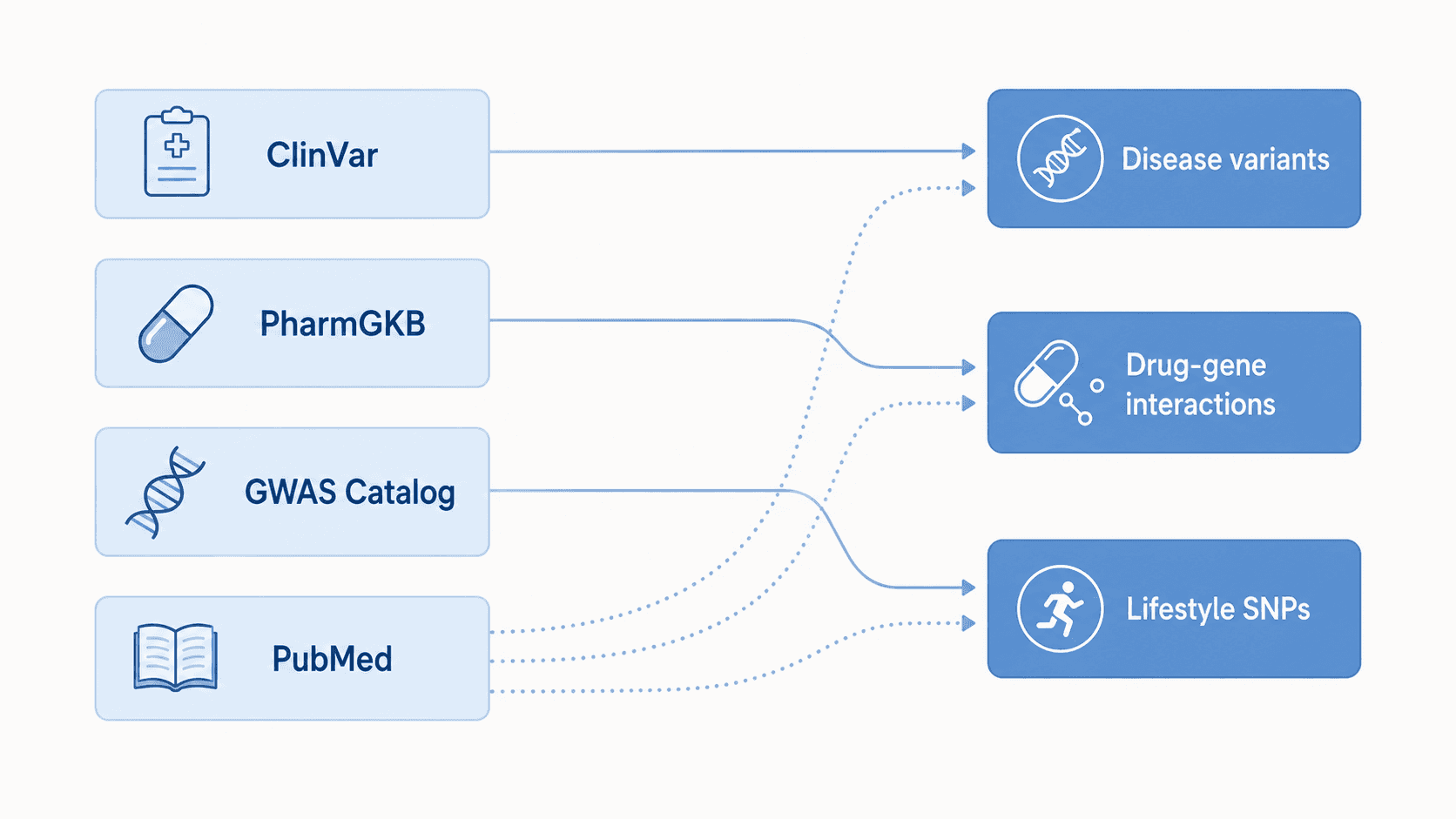
For YMYL content, source quality is more important than volume. GenoSight's insights come from four primary databases: ClinVar for disease-associated variants, the GWAS Catalog for trait associations, PharmGKB for drug-gene interactions, and PubMed for the underlying primary literature. This page explains what each source is, what evidence tier we use from it (ClinVar gold-star, PharmGKB Levels 1A–2B), and why we chose primary sources over aggregator sites or wiki-style references.
Why source quality matters more than source volume
A genetic report can cite hundreds of papers, or it can cite four databases — and in many cases the four-database version is more trustworthy. The reason is that primary sources have curation discipline that aggregator sites don't.
Aggregator sites and wikis (the most common example: SNPedia) compile findings across many papers without the same gatekeeping. They're useful for breadth but they include weak findings alongside strong ones. A reader has to do the evidence triage themselves — and most readers can't.
Primary databases curate by design. ClinVar uses a star-based confidence system. PharmGKB grades evidence on a published rubric (1A through 4). The GWAS Catalog reports significance thresholds and study sizes. When a finding makes it into one of these databases at the tier we use, that's already meaningful.
GenoSight's choice: cite primary sources directly, surface the evidence tier alongside every finding, and let the reader see exactly what's behind a recommendation. Source choices like this also bound what's possible to claim — for the broader picture of what consumer-array analysis can and can't catch, the source list matters as much as the analysis pipeline.
ClinVar — disease-associated variants
ClinVar is the public archive of variants of clinical significance, maintained by the National Center for Biotechnology Information. Each variant entry includes:
- The clinical interpretation (pathogenic, likely pathogenic, uncertain significance, likely benign, benign)
- The submitting laboratories (often multiple, sometimes with conflicting interpretations)
- A gold-star confidence rating based on the level of review:
- 0 stars: no assertion criteria provided
- 1 star: criteria provided, single submitter
- 2 stars: criteria provided, multiple submitters with no conflicts
- 3 stars: reviewed by an expert panel
- 4 stars: practice guideline
GenoSight scans ClinVar variants against your raw file with gold-star filtering — we surface variants at higher confidence tiers and flag lower-confidence findings explicitly when they're surfaced at all. As of the current build, this covers 341,375 ClinVar SNPs (true single-nucleotide variants only — indels are excluded because consumer arrays don't reliably call them).
PharmGKB — drug-gene interactions
PharmGKB is the Pharmacogenomics Knowledgebase, maintained at Stanford. It catalogs drug-gene interactions and grades the evidence for each on a published rubric:
| Level | Meaning |
|---|---|
| 1A | Variant–drug pair with consensus from multiple major studies and clinical guideline support |
| 1B | Strong evidence from multiple studies; some clinical implementation |
| 2A | Moderate evidence from multiple studies in different cohorts |
| 2B | Moderate evidence; replicated but smaller cohorts |
| 3 | Single significant study or non-replicated evidence |
| 4 | Case reports or in-vitro evidence only |
GenoSight surfaces only PharmGKB Levels 1A through 2B in reports — the tiers where evidence is strong enough to inform real prescribing decisions. Levels 3 and 4 are excluded by default because the evidence isn't yet strong enough to act on; surfacing them risks overconfident recommendations.
When a PGx finding appears in your report, the cited evidence level is shown alongside it. That tells you and your clinician exactly how confident the finding is.
GWAS Catalog — trait associations
The GWAS Catalog is the European Bioinformatics Institute's curated catalog of genome-wide association studies. Each entry records the trait studied, the variants associated, the effect size, the p-value, and the cohort sizes.
GenoSight uses GWAS Catalog evidence to support lifestyle SNP findings — particularly for variants where individual study evidence accumulates across multiple cohorts. The catalog's transparency about cohort size and significance thresholds is what makes it useful for our purpose; we can show the reader (and their clinician) the underlying study basis for a given trait association.
For lifestyle SNP recommendations, GWAS Catalog evidence is reported alongside the catalog entry rather than synthesized into vague aggregate claims.
PubMed — the underlying primary literature
PubMed is the National Library of Medicine's database of biomedical publications. Where the three databases above curate findings, PubMed provides the primary literature behind them.
GenoSight uses PubMed citations to anchor specific claims in the underlying studies. When a report links a finding to a specific paper — the way the methylation example article links MTHFR C677T to Frosst et al. 1995, or homocysteine reference ranges to a MedlinePlus reference — the reader can verify the source directly.
This is the difference between citing "studies suggest" and citing a specific paper a clinician can read in five minutes.
What we don't use, and why
A few sources that are common in the consumer-genomics interpretation space, that GenoSight does not use:
- SNPedia is a wiki of variant interpretations. Useful for breadth, but its content license (CC-BY-NC-SA) prohibits commercial use, and the wiki model includes findings of varying evidence quality without consistent curation. Other tools (notably Promethease) lean heavily on SNPedia; we don't.
- Genetic Lifehacks article prose. The site has excellent editorial depth and we use its 19-topic taxonomy as a target for our own content clusters — but the prose is copyrighted and we don't paraphrase it. Our content is sourced from the four primary databases above.
- Aggregator and content-farm sites. Recommendations sourced from broad aggregator sites without primary citations don't make it into reports.
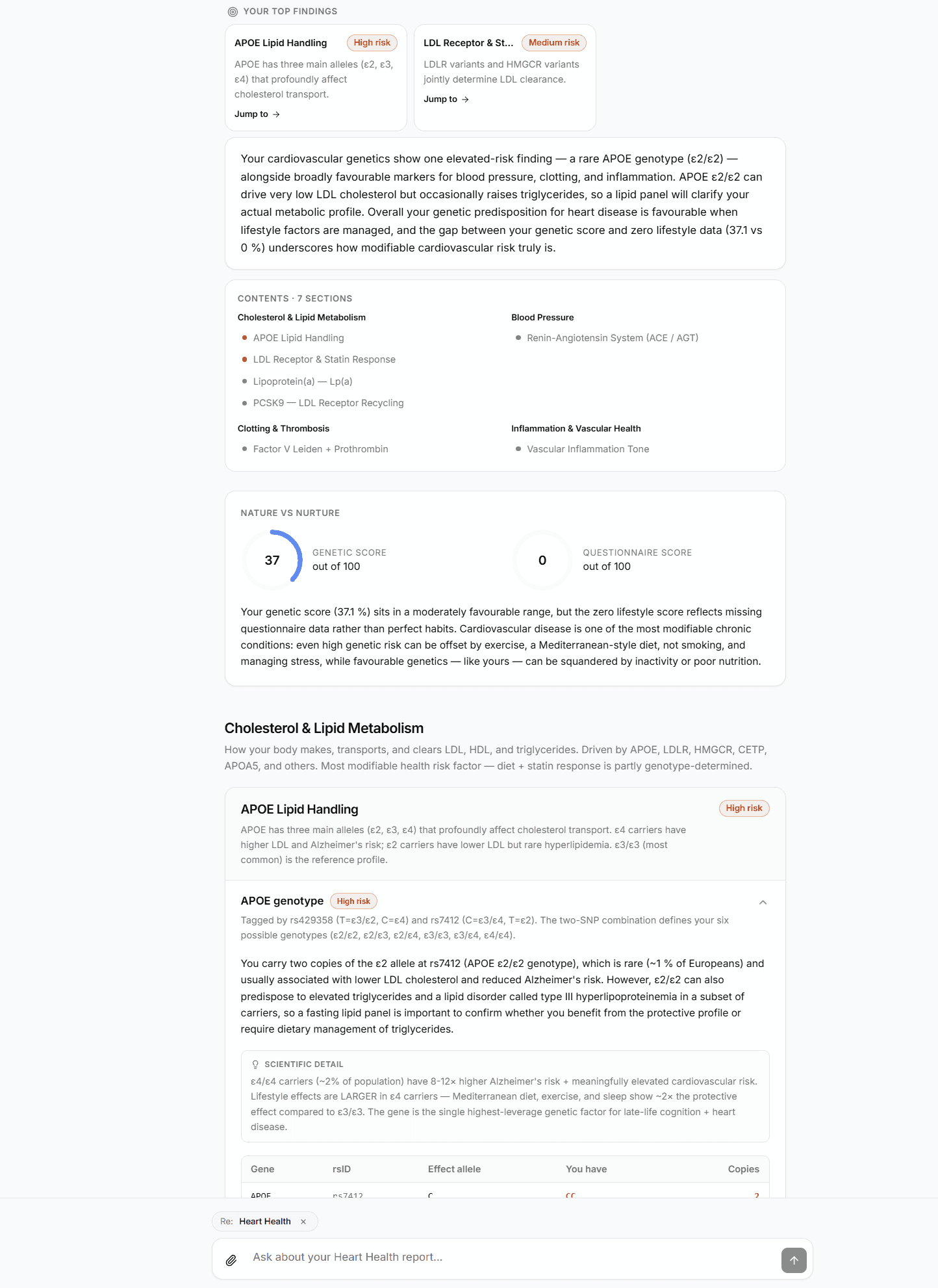
How citations show up in your report
Every claim in your GenoSight report has a citation chain you can inspect:
- Variant-level findings cite the relevant database entry directly (ClinVar, PharmGKB, or GWAS Catalog)
- Synthesis-level claims (interactions between findings, recommendations) cite the underlying findings they're built on
- Specific clinical claims with primary literature link to PubMed entries
If you ask a follow-up question in the chat, the answer carries the same citation discipline — you can ask "what's the evidence behind that recommendation?" and get the source spelled out.
Why this matters for YMYL content
Google's quality raters have explicit guidance for "Your Money or Your Life" content — anything affecting health, finance, or major life decisions — that prioritizes E-E-A-T (Experience, Expertise, Authoritativeness, Trust). Source quality is a load-bearing component.
For us, primary-source citation isn't just an SEO concern. It's how we earn the right to be in this space. A health recommendation backed by a verifiable ClinVar gold-star entry is something a reader can take to their doctor. A recommendation backed by "studies suggest" can't be defended in the same way.
This citation discipline also runs into hard limits about what consumer arrays can detect — being honest about both ends matters.
See the citations behind your report
Every finding includes its underlying source and evidence tier. Free trial, 250 credits.
Key takeaways
- For YMYL content, source quality matters more than source volume. Primary databases have curation discipline that aggregator sites don't.
- GenoSight uses four primary sources: ClinVar (disease variants), PharmGKB (drug-gene interactions), GWAS Catalog (trait associations), and PubMed (primary literature).
- Evidence tiering is explicit: ClinVar gold-star filtering for disease variants, PharmGKB Levels 1A–2B for drug-gene interactions.
- We don't use SNPedia (license + curation issues) or paraphrase Genetic Lifehacks' content (copyrighted; taxonomy used as a content target only).
- Every claim in a GenoSight report has a citation chain you can inspect — source, evidence tier, and (where relevant) the underlying paper on PubMed.